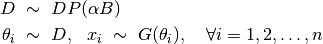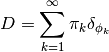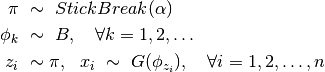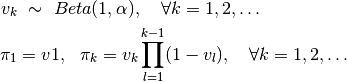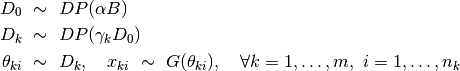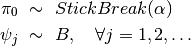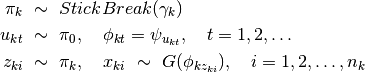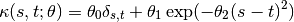Nonparametric Models¶
Bayesian nonparametric models, such as DP mixture models, provide a flexible approach in which model structure (e.g. the number of components) can be adapted to data. The capability and effectiveness of such models have been proven in many applications.
This flexibility, however, leads to new questions to probabilistic programming – how to express models whose size/structure can vary. The Julia macro system makes it possible to address this problem in an elegant way due to is lazy evaluation nature.
Dirichlet Process Mixture Model¶
Dirichlet process mixture model (DPMM) is one of the most widely used in Bayesian nonparametrics. The formulation of DPMM is given by
Here, 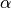 is the concentration paramater,
is the concentration paramater, 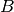 is the base measure, and
is the base measure, and 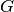 denotes the component models that generate the observed samples. The model specification for a DPMM is given as below. It has been shown that
denotes the component models that generate the observed samples. The model specification for a DPMM is given as below. It has been shown that 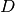 is almost surely discrete, and can be expressed in the followin form:
is almost surely discrete, and can be expressed in the followin form:
Hence, there exists positive probability that some of the components will be repeatedly sampled, and thus the number of distinct components is usually smaller than the number of samples. In practice, it is useful to construct a pool of components 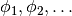 , and introduce an indicator
, and introduce an indicator 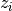 for each sample
for each sample 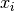 , such that
, such that 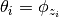 . To make this explicit, we can reformulate the model as below
. To make this explicit, we can reformulate the model as below
Here, \pi, a sample from the stick breaking process, is an infinite sequence that sums to unity (i.e. 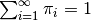 ). The values of
). The values of 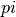 are defined as
are defined as
The model specification for this formulation is given below
@model DPMixtureModel{B, G} begin @constant n::Int # the number of observed samples @hyperparam alpha::Float64 # the concentration parameter pi ~ StickBreak(alpha) for k = 1 : Inf phi[k] ~ B end # samples for i = 1 : n z[i] ~ pi x[i] ~ G(phi[z[i]]) end end
Remarks:
- The parametric setting makes it possible to use arbitrary base distribution
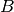 and component
and component 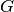 here, with the same generic formulation.
here, with the same generic formulation. - In theory,
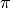 is an infinite sequence, and therefore it is not feasible to completely instantiate a sample path of
is an infinite sequence, and therefore it is not feasible to completely instantiate a sample path of 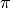 in computer. This variable may be marginalized out in the inference, and thus directly querying
in computer. This variable may be marginalized out in the inference, and thus directly querying 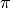 is not allowed.
is not allowed. 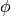 has infinitely many components. However, only a finite number of them are needed in inference. The compiler should generate a lazy data structure that only memorizes the subset of components needed during the inference. In particular,
has infinitely many components. However, only a finite number of them are needed in inference. The compiler should generate a lazy data structure that only memorizes the subset of components needed during the inference. In particular, phi[k]is constructed and memorized when there is anisuch thatk = z[i]. Some efforts (e.g. the LazySequences.jl) have demonstrated that lazy data structure can be implemented efficiently in Julia.
Hierarchical Dirichlet Processes¶
HDP is an extension of the DP mixture models, which allows groups of data to be modeled by different DPs that share components. The formulation of HDP is given below
Using 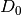 as a base measure for the DP associated with each group, all groups share components in
as a base measure for the DP associated with each group, all groups share components in 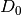 while allowing potentially infinite number of components. This formulation can be re-written (equivalently) using Pitman-Yor process, as follows
while allowing potentially infinite number of components. This formulation can be re-written (equivalently) using Pitman-Yor process, as follows
Then for each 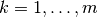 ,
,
Here is a brief description of this procedure:
- To generate
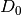 , we first draw an infinite multinomial distribution
, we first draw an infinite multinomial distribution 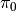 from a Pitman-Yor process with concentration parameter
from a Pitman-Yor process with concentration parameter 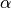 , and draw each component
, and draw each component 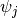 from
from 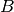 . Then
. Then 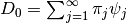 .
. - Then for each group (say the k-th one), we draw
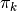 from a stick breaking process and draw each component from
from a stick breaking process and draw each component from 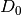 . Note that drawing a component
. Note that drawing a component 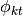 from
from 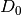 is equivalent to choosing one of the atoms in
is equivalent to choosing one of the atoms in 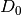 , which can be done in two steps: draw
, which can be done in two steps: draw 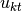 from
from 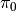 and then set
and then set 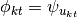 . In other words, the
. In other words, the 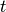 -th component in the
-th component in the 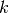 -th group is identical to the
-th group is identical to the 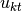 -th component in
-th component in 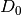 .
. - Finally, to generate the
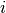 -th sample in the
-th sample in the 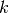 -th group, denoted by
-th group, denoted by 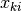 , we first draw
, we first draw 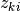 from
from 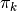 and use the corresponding component
and use the corresponding component 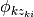 to generate the sample.
to generate the sample.
This formulation can be expressed using the DSL as below:
Gaussian Processes¶
Gaussian process (GP) is another important stochastic process that is widely used in Bayesian modeling. Formally, a Gaussian process is defined to be a a function-valued distribution 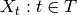 , where
, where 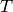 can be arbitrary domain, such that any finite subset of values in
can be arbitrary domain, such that any finite subset of values in 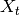 is normally distributed. A Gaussian process is characterized by a mean function
is normally distributed. A Gaussian process is characterized by a mean function 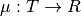 and a positive definite covariance function
and a positive definite covariance function 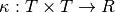 . The covariance function is typically given in a parametric form. The following is one that is widely used
. The covariance function is typically given in a parametric form. The following is one that is widely used
In many application, the GP is considered to be hidden, and observations are a noisy transformation of the samples generated from the GP, as
The following model specification describes this model.
The GaussianProcess distribution here is a high-order stochastic function, which takes into two function arguments and generates another function. This is readily implementable in Julia, where functions are first-class citizens like in many functional programming languages.